|
|
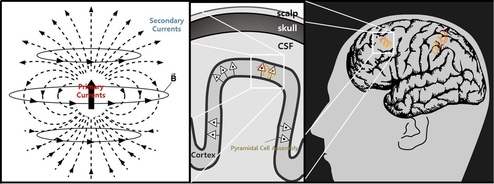
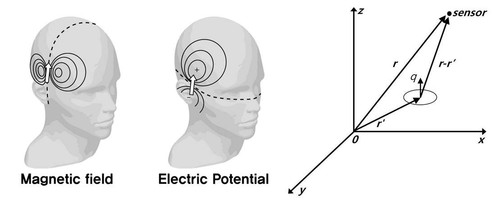
Electroencephalography (EEG) Source Localization EEG and MEG are the only techniques among current methods which uniquely provide a temporal resolution under 100ms. They provide superior temporal resolution over the other imaging technologies such as functional magnetic resonance imaging (fMRI) and positron emission tomography (PET). However, they suffer from the poor spatial resolution since their signals are distorted by many physical and physiological factors. As a consequence, it is hard to use EEG and MEG in practical situation for a brain function imaging.
Localization of brain signal sources from EEG/MEG has been an active area of research. Currently, there exists a variety of approaches such as MUSIC, M-SBL, and etc. These algorithms have been applied for various clinical examples and demonstrated excellent performances. However, when the unknown sources are highly correlated, the conventional algorithms often exhibit spurious reconstructions. To address the problem, we proposed a new algorithm that generalizes M-SBL by exploiting the fundamental subspace geometry in the multiple measurement problem (MMV) [1]. Experimental results using simulation and real phantom data show that the proposed algorithm outperforms the existing methods even under a highly correlated source condition [2].
Electroencephalography (EEG) for Brain Network Imaging and Disease Diagnosis EEG has been used as a common method to study brain function and disorder with high temporal resolution. Still, unlike fMRI, network analysis using EEG has been poor due to its limited spatial resolution. However, recent findings in EEG microstates identified that it is possible to see the brain network which coincides with fMRI results by using EEG topography. This study implies that if we interpret EEG with carefully chosen network method, we can obtain DMN results with rich temporal information while the general trend of network still remains comparable to fMRI.
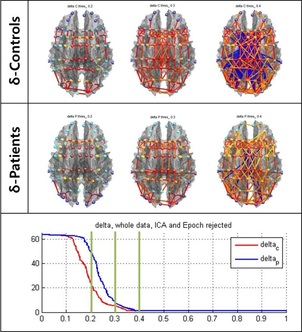
Brain network is usually represented as a graph consisting of nodes and edges. Connectivity matrix is constructed by thresholding it. However, currently there is no agreed-upon method for choosing an optimal threshold. Moreover, determining a fixed threshold will cause very different networks, even though they are originated from a same data. This can incur not only different interpretations but also losing the wealth of information by discarding the data below the threshold. Here, we employed a threshold-free network analysis based on persistent homology framework. This method observes the topological changes of the networks generated over every possible threshold. Thus, our applied method enables us to quantify persistent variations of topological features in brain network. We examined the resting state EEG data, and our results revealed that the evolving topological features, especially persistence landscape, distinguished the MDD patients from normal controls with statistically significant differences. Near Infrared Spectroscopy (NIRS) for Brain Network Imaging and Disease Diagnosis Even though making a quick and accurate diagnosis for cerebrovascular diseases such as cerebral hemorrhage and infarction is critical in acute state, it is hard to shorten the time of current system using CT or MRI. Moreover, there is a growing need to continuously image a premature baby’s brain oxygen level to prevent the disease such as cerebral palsy. Therefore, in this research we aim to develop quantitative indicator and statistical analysis tool for these situations and reconstruct a high resolution brain activity image using functional near infrared spectroscopy (fNIRS). Final goal of our research is to obtain a portable fNIRS brain imaging technology which can provide the continuous real time image in emergency situation.
Reference
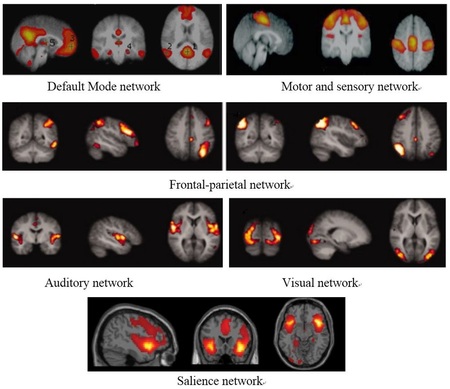
[1] Jong Chul Ye, Jong Min Kim, and Yoram Bresler, " Subspace penalized sparse learning for joint sparse recovery", IEEE International Conference on Acoustics, Speech and Signal Processing(ICASSP), Vancouver, Canada, May 2013 [2] Jaejun Yoo, Jong Min Kim, Chang Hwan Im, and Jong Chul Ye, "Neuroelectromagnetic Imaging of Correlated Sources Using a Novel Subspace Penalized Sparse Learning", IEEE International Symposium on Biomedical Imaging (ISBI), California, San Francisco, USA, April 2013 Resting State Functional Magnetic Resonance Imaging Recently, the state of the brain networks while at rest has been the subject of many studies and it seems to play a significant role in the study of several brain disorders. It is the important concept of functional connectivity between certain brain regions that gives rise to the formation of resting state networks. Resting state functional connectivity refers to the connectivity between different brain regions in terms of their functionality assessed during resting conditions. It reflects the patterns of statistical dependencies between distributed neural elements and is often expressed in terms of the correlation between regional time series. Several studies have been investigating the existence of different resting state networks corresponding to different sets of functionally connected brain regions.
Functional Magnetic Resonance Imaging (fMRI) is a new imaging technique that uses the physical phenomenon of nuclear magnetic resonance and the associated technology of magnetic resonance imaging to map human brain function by measuring neural activity through the detection of changes in blood flow. Primarily, fMRI uses the blood-oxygen-level-dependent (BOLD) signal to reflect spatially localized changes in blood oxygenation and brain hemodynamics in response to local neural activity. In more details, BOLD fMRI is sensitive to the local changes in deoxygenated hemoglobin due to neural activation. The hemoglobin molecule has magnetic properties that changes depending on whether or not it is bound to oxygen. If it is bound to oxygen, forming oxygenated hemoglobin, it will be diamagnetic which means it will have zero magnetic moment and therefore will not have any magnetic property. However when it is not bound to oxygen, called deoxygenated hemoglobin, it becomes paramagnetic, having significant magnetic moment that can affect the magnetization in the MRI machine.
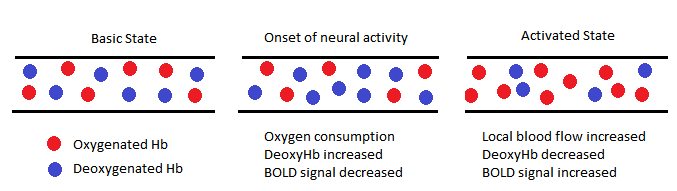
When a neuron is activated, it draws oxygen from the blood stream, leading to an increase in the concentration of deoxy-hemoglobin in the blood. After the onset of neuron activity, the active neurons consume the oxygen available in the local blood stream and thus the relative level of deoxy-hemoglobin increases more which leads to an initial decrease in BOLD fMRI signal. After this initial increase in deoxy-hemoglobin concentration, local blood flow increases in response to neuronal activity, providing a large amount of oxygen-rich blood, more than the required amount to preserve oxygen consumption due to neural activity. This in turn results in local decrease in the concentration of deoxy-hemoglobin. Considering the fact that deoxy-hemoglobin is paramagnetic, a reduction in its concentration results in an increase in the homogeneity of the static magnetic field which yields an increase in BOLD fMRI signal. At last, the concentration of deoxy-hemoglobin slowly returns to its normal level and the BOLD fMRI signal decays till reaching its original baseline level. In recent years there has been great interest in the application of fMRI in cases when the subject is at rest and does not perform any task. Currently known as resting state fMRI or functional connectivity MRI, it is a powerful method of fMRI that can evaluate state of the brain and its resting state networks when the subject is at rest by means of measuring the low frequency fluctuations in the BOLD signal. The range of these frequency fluctuations is between 0.01 and 0.1 Hz. It has been suggested that these signal variations are of neuronal origin that characterize the neural baseline activity of brain and can correspond to the functional resting state networks as well. Since it examines the temporal correlations between segregated brain regions, resting state fMRI has made a considerable breakthrough in the functional connectivity studies as well as the identification of resting state networks in the brain.
Over the past few years, researchers have applied resting state fMRI technique to discover the resting state networks alterations in different brain disorders in order to gain a better understanding of the progression of the disease, consequent symptoms, possible causes as well as possible treatments. The utilization of resting state fMRI on Parkinson’s and Alzheimer’s disease patients forms the basis of our research. To analyze the networks, the conventional methods of seed-based correlation analysis and independent component analysis are employed using SPM (Statistical Parametric Mapping) and FSL (fMRI Software Library) toolboxes, respectively. Moreover, conventionally the brain network connectivity has been thresholded based on a fixed threshold value; however, in our recent research, the novel method of persistent homology analysis is further applied to study the changes of the networks over a range of threshold values. Furthermore, the method examines the underlying topological changes of the network, defined by the changes in the number of connected components as well as holes, as the threshold value changes. We performed persistent homology analysis on our resting-state fMRI data and observed significant differences in patients’ group compared to normal subjects, quantified through the statistical analysis of persistence landscapes. |
|
ABOUT US
Our research activities are primarily focused on the signal processing and machine learning for high-resolution high-sensitivity image reconstruction from real world bio-medical imaging systems. Such problems pose interesting challenges that often lead to investigations of fundamental problems in various branches of physics, mathematics, signal processing, biology, and medicine. While most of the biomedical imaging researchers are interested in addressing this problem using off-the-self tools from signal processing, machine learning, statistics, and optimization and combining their domain-specific knowledge, our approaches are unique in the sense that I believe that actual bio-medical imaging applications are a source of endless inspiration for new mathematical theories and we are eager to solve both specific applications and application-inspired fundamental theoretical problems.
|
CONTACT US
Bio Imaging. Signal Processing & Learning
Graduate School of AI KAIST 291 Daehak-ro, Yuseong-gu Daejeon 305-701, Korea Copyright (c) 2014, BISPL All Rights Reserved. |
|